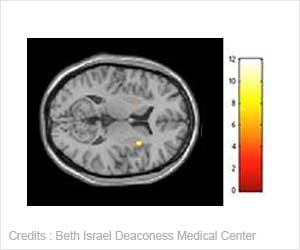
Researchers have analysed immunocompromised patient blood samples and shown HCMV is not mutating any more than other DNA viruses.
TOP INSIGHT
HCMV mutation rates are similar to other DNA viruses, providing some reassurance for the development of vaccines, as unlike the hyperdiverse RNA viruses, HCMV is unlikely to mutate to evade vaccines.
Now using new viral genome sequencing, bioinformatic and modelling tools developed at UCL, researchers have analysed immunocompromised patient blood samples and shown HCMV is not mutating any more than other DNA viruses. Instead, the frequent occurrence of mixed-infections caused by genetically different HCMV strains leads to findings of high genome diversity within an individual.
The study, funded by the European Union, has been published in Proceedings of the National Academy of Sciences. Lead author, Professor Judith Breuer (UCL Division on Infection & Immunity), said: "It has been claimed that HCMV is as diverse as the more error prone RNA viruses such as HIV and Hepatitis C Virus and this has led to a lot of confusion in the field.
"Using our novel genome sequencing and bioinformatics modelling approaches, we have shown that the apparently high HCMV diversity seen in some clinical samples is caused not by HCMV mutation rather it is due to frequent co-infection with multiple strains of HCMV.
New UCL sequencing tool
The Breuer Lab has pioneered whole pathogen genome sequencing, by methods called 'targeted enrichment', to generate high quality whole viral genomes directly from clinical material, without the need for viral culture or polymerase chain reaction (PCR). She has combined these methods with cutting-edge bioinformatics including the newly developed Haplotype Reconstruction of Longitudinal Sequence Data (HaRoLD).
"This process is comparable to sorting several jigsaw puzzles that have been jumbled up together," added Professor Breuer.
"The new bioinformatics methods for re-constructing individual genome sequences from a mixed infection will allow us to identify the factors, including recombination, that drive HCMV evolution in a patient, providing a precision approach to managing infections."
Source-Eurekalert
 MEDINDIA
MEDINDIA



 Email
Email