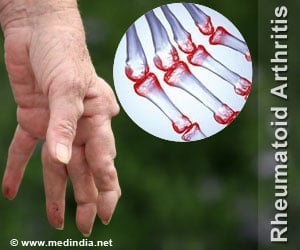
Distinct gene expression signatures in rheumatoid arthritis patients could help predict how these individuals will respond to tumor necrosis factor inhibitors.
‘Distinct gene expression signatures in rheumatoid arthritis patients could help predict how these individuals will respond to tumor necrosis factor inhibitors.’





Distinct gene expression signatures in rheumatoid arthritis patients
could help rheumatologists predict how these individuals will respond to
tumor necrosis factor inhibitors, and may one day enable a more
personalized approach to RA therapy, revealed new research findings
presented this week at the 2016 ACR/ARHP Annual Meeting in Washington."The aim of this study was to better understand the impact of the type-I interferon ratio that predicts non-response to TNFi therapy on a major inflammatory cell type involved in RA," said Theresa L. Wampler Muskardin,a rheumatologist in the Department of Pediatric and Adolescent Medicine, and a lead author of the study. "Effects of type-I interferon on single cells and immune cell subtypes may be missed when we analyze whole blood or mixed cell populations. Using single cell gene expression technology, we hoped to find differences in expression of select genes between treatment responders and non-responders, which could ultimately lead to a blood test that we can use to guide treatment decisions in RA patients prior to starting biologic therapy."
The researchers used single cell expression analysis to study whether monocyte gene expression was significantly different among RA patients based on their pre-treatment blood serum IFN- β/α ratio. They isolated single classical (CL) and single non-classical (NC) blood-derived monocytes from 15 seropositive RA patients before TNFi therapy. The patients were divided into two groups according to their pre-TNFi serum ratio: six patients in IFN-β/α>1.3 and nine patients in IFN-β/α<1.3. The researchers performed unsupervised hierarchical clustering of 87 target genes on the single monocytes.
Results of the testing showed that JAK1 and IL1A strongly differentiated between the two groups. In NC cells only, expression of STAT2, ILT7, PKR, TLR7 and IRAK1 were more likely in the non-response patient group. In CL cells only, expression of IFIT2 and CD36 was more likely. In a multivariate logistic regression, IL1A, CD32a, IL-8, TYK2 and IRAK1 in NC plus CL monocytes aligned with the treatment response groups. In comparison with that mixed monocyte model, IL-8 and IRAK1 in NC, and CXCR3 in CL showed even stronger alignment with the treatment response groups. STAT2 strongly predicted the response group in NC cells alone. CXCL9 strongly predicted the response group in CL cells alone.
Their previous work showed that an IFN- β/α ratio>1.3 is predictive of non-response to anti-TNF therapy in RA patients. In this study, their findings show that gene expression in monocyte subsets differ in RA patients who have an IFN-β/α ratio>1.3, the ratio of type-I IFNs that predicts non-response to anti-TNF therapy.
Advertisement
Future studies should focus on monocyte subsets to identify possible molecular differences that could also determine treatment response to TNFi biologics. In the future, rheumatologists may be able to tailor therapy to an RA patient based on his or her underlying disease biology, Dr. Muskardin concluded.
Advertisement