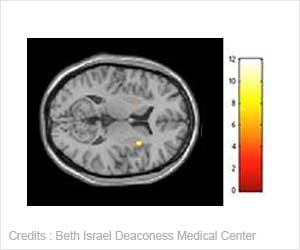
Researchers have combined experimental data from X-ray crystallography, NMR spectroscopy, cryoelectron microscopy and lipidomics and have built a 'virtual virus'.

The computer simulation begins by rendering the virtual virus as a relatively large, 73-nanometer ball of loosely packed lipids, which then relaxes down into a smaller, 59-nanometer virion within 300 nanoseconds, an imperceptible amount of time on the macroscopic level, but roughly 1/15th of the simulation's total run time. After this the viral spike proteins are embedded into the lipid envelope individually, before adding solvent to the system.
From the simulation, researchers found that the viral spike proteins protruding from the virion's membrane spread out, rather than aggregating close together. This was the key to the strength of the interactions between influenza A virions and host cells, which are determined by the number of spike proteins that can engage with receptors.
With this understanding of the membrane envelope's structural dynamics, scientists have got an insight into the wide-ranging survival times of the virion in different environments, such as fresh-water rivers.
Source-Medindia
 MEDINDIA
MEDINDIA



 Email
Email