Scientists at Yale University say that the origin and evolution of the placenta and uterus in mammals is associated with evolutionary changes in a single regulatory protein.
Scientists at Yale University say that the origin and evolution of the placenta and uterus in mammals is associated with evolutionary changes in a single regulatory protein.
"Many past studies have shown that genes are regulated and altered by changes within their own structures. This is the first work suggesting that the evolution of transcription factors - separate regulatory proteins - may play an active role in the origin and evolution of structural innovations like the placenta and uterus," said senior author Gunter Wagner, the Alison Richard Professor of Ecology and Evolutionary Biology at Yale.In their study report, published in the journal Proceedings of the National Academy of Sciences, the researchers describe pregnancy as a biologically unusual situation where one organism lives and develops inside another that is genetically different.
Ordinarily, the immune system identifies and destroys the dissimilar tissue as if it were a parasite, but in some early mammals, changes 'turned down' the immune system, allowing the developing embryo to grow and thrive unchallenged by the maternal immune response.
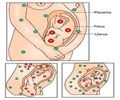
The evolution of the uterus and placenta made it possible for mammals to protect their growing young, and to ensure they were not exposed to an unpredictable environment, like their egg-laying relatives.
The Yale researchers say that their study has identified one of the genetic switches that tempered the immune system to enable the formation of the placenta and internal development of young.
The team said that they analysed DNA from many species of mammals, including resurrecting genes from the extinct ancestors of mammals, and found that a crucial regulatory link in the evolution of pregnancy involved the altered function of a transcription factor protein, HoxA-11.
It, however, does not exist in marsupials like opossums or wallabies, where there is a brief and rudimentary pregnancy followed by development of the offspring outside the mother. Egg-laying mammals, such as the platypus, also lacked it.
Graduate student Vincent Lynch, lead author of the study, said: "We are writing a different chapter. In this case the function of a major regulatory tool was altered - it is like we found a redesigned hammer."
Source-ANI
SRM
 MEDINDIA
MEDINDIA


 Email
Email