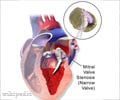
Penn team used the new gene-editing technique to rapidly generate induced pluripotent stem cells containing the patient's specific TNNT2 gene variant.
TOP INSIGHT
The genetic avatars containing the patient's unique TNNT2 variant was then induced to mature into heart muscle cells to test their responses to the adrenaline-like chemical.
A team led by Kiran Musunuru, MD, PhD, an associate professor of Cardiovascular Medicine and Genetics, and Anjali Tiku Owens, MD, an assistant professor of Cardiovascular Medicine, needed to find out whether the variant could explain her heart disease and whether the patient's two children and six grandchildren should also undergo genetic screening. To do so, the team used a stem-cell-based assay they were developing to provide highly personalized results for the patient and her family. The Penn team's findings are described this week in Circulation.
"More and more, through sequencing done for medical reasons or done by direct-to-consumer companies, people learn they have variants of disease genes with uncertain significance to their health," said Owens. "With our assay, we now have a way to start determining the potential impact of these variants on the health of patients and their family members."
In the 10 weeks between the patient's first and second visits to the clinic in 2017, the Penn team used the new gene-editing technique developed in Musunuru's lab to rapidly generate induced pluripotent stem cells containing the patient's specific TNNT2 gene variant. These tailored stem cells--personalized avatars, as it were--showed a normal response to a chemical that induces faster beating in heart cells, suggesting the variant was not disease-causing. Guided by this finding, the team recommended that the patient's family not undergo genetic screening, but still be monitored in the future for possible thickening of the heart wall.
"We believe this is the first time that stem cells have been used to impact patient and family care for cardiovascular disease," said Musunuru. As part of the development of the test the team created a library of TNNT2 gene variants and isolated 14 unique types. Most of these were classified as pathogenic, or disease-causing, according to the new assay. The platform allows for rapid insertion of specific gene variants into stem cells using a combination of CRISPR and other gene-editing tools. Results showed that the TNNT2 variant from Owens' patient was benign.
"This could help any person in the world found to have a variant in TNNT2 make better choices about their care," says Musunuru.
 MEDINDIA
MEDINDIA


 Email
Email