A sophisticated procedure to track cell growth in yeast at very low nutrient concentrations was devised by scientists.
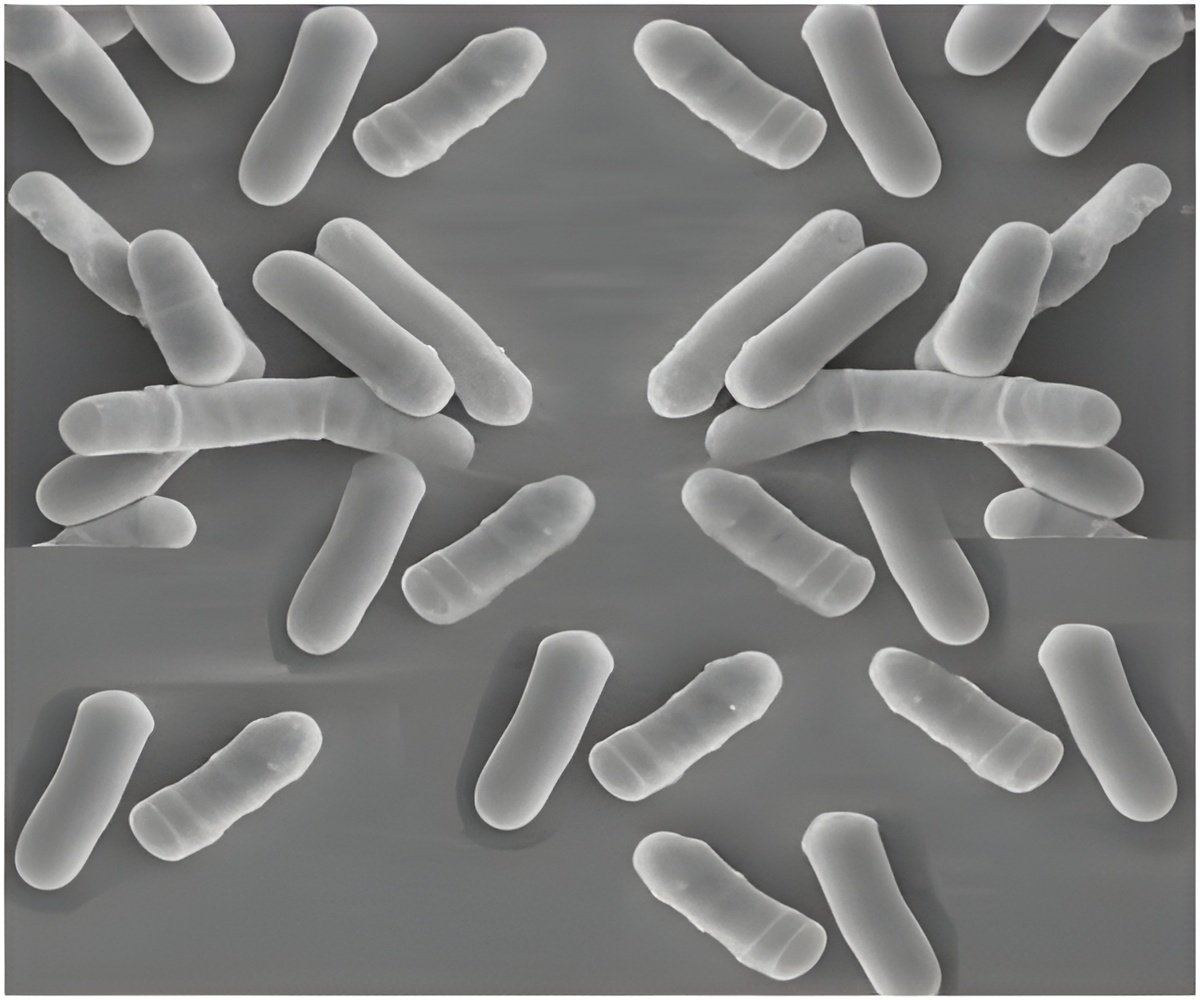
The researchers studied growth rates and lag times in both lab strains and wild yeast by varying the amount of its prime carbon food source, glucose.
They confirmed the prediction made over 60 years ago by Noble-prize-winning biologist Jacques Monod regarding changes in microbial growth rates with limited nutrients (the Monod equation).
They also found significant differences among strains in both the average lag response (the amount of time it takes to transition from cell quiescence to restarting cell growth) and average growth rates in response to different environmental conditions.
Lead author, Naomi Ziv said that the different strain variances that are seen suggest that the extent of nongenetic heterogeneity is itself genetically determined.
Source-ANI
 MEDINDIA
MEDINDIA



 Email
Email