Researchers developed a way to identify the carcinogenic cells in the liver, a pattern of mutations that serve as a fingerprint for aflatoxin.
TOP INSIGHT
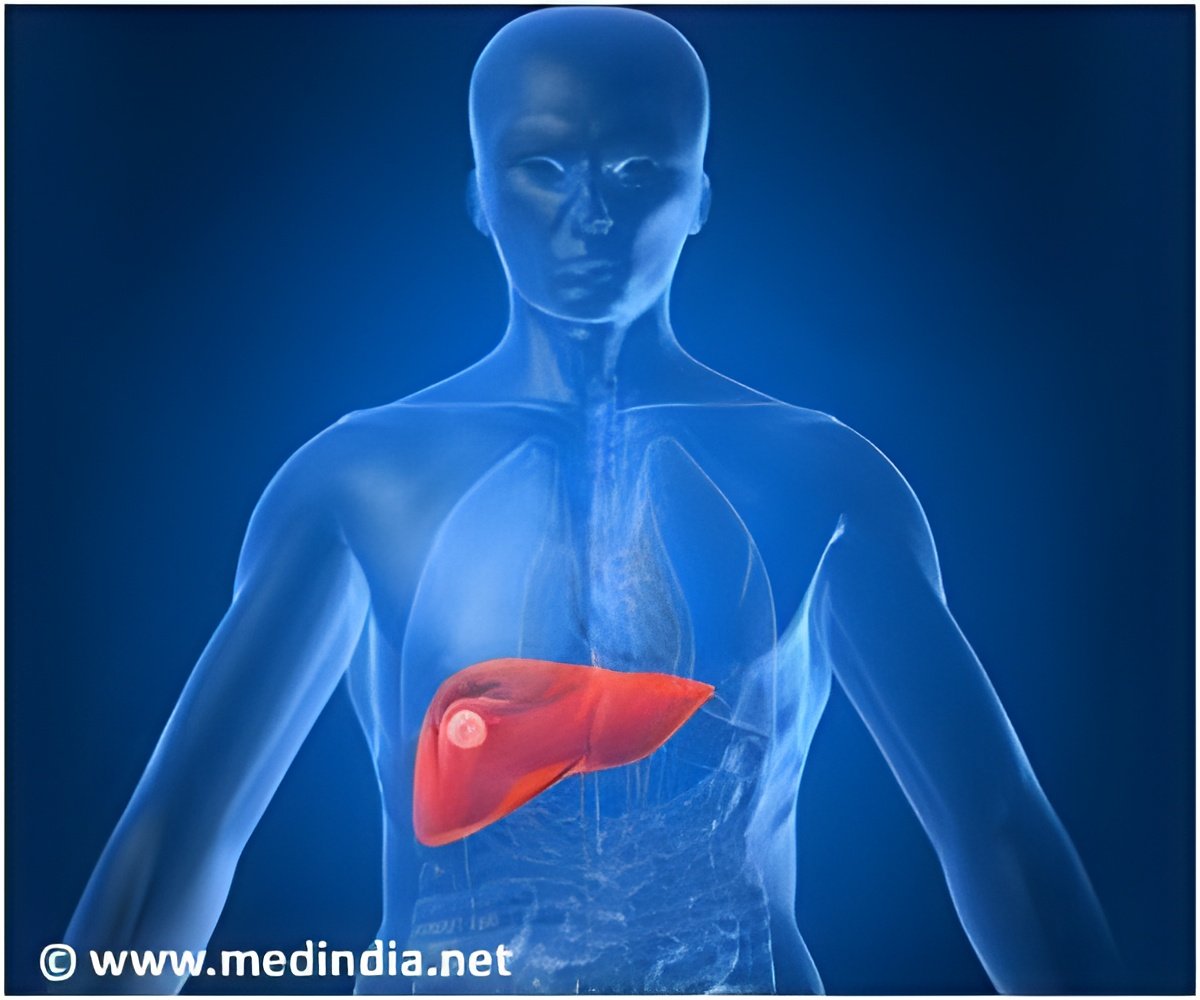
Exposure to a fungal product called aflatoxin is believed to cause 80 percent of liver cancer cases. The technique of DNA sequencing enables the cancer to be detected early.
This fungus is often found in corn, peanuts, and other crops that are dietary staples in those regions.
The new technique can determine whether liver cells have been exposed to aflatoxin and help predict whether someone has a high risk of developing liver cancer, potentially many years before tumors actually appear.
"What we're doing is creating a fingerprint," said John Essigmann, Professor at MIT.
"It's really a measure of prior exposure to something that causes cancer," added Essigmann, who is the senior author of a paper describing the findings in the Journal Proceedings of the National Academy of Sciences.
In the new study, the MIT team set out to see if they could identify mutations produced by aflatoxin long before cancer develops.
The researchers sequenced DNA from those tumors and also removed from liver cells; only 10 weeks after exposure, before tumors developed.
To find mutations at 10 weeks, the researchers used a powerful genome sequencing technique that can identify very rare mutations which occur in about one in 10 million to 100 million DNA base pairs.
Unlike most DNA sequencing techniques, the one used in this paper, developed by researchers at the University of Washington, combines data from two complementary strands of DNA.
With the new technique, the researchers found that at 10 weeks, a distinctive pattern of mutations that can serve as a "fingerprint" for aflatoxin exposure had already emerged.
Source-IANS
 MEDINDIA
MEDINDIA
 Email
Email