Except an association with the human papillomavirus, no validated molecular profile of head and neck squamous cell carcinoma (HNSCC) has been established.
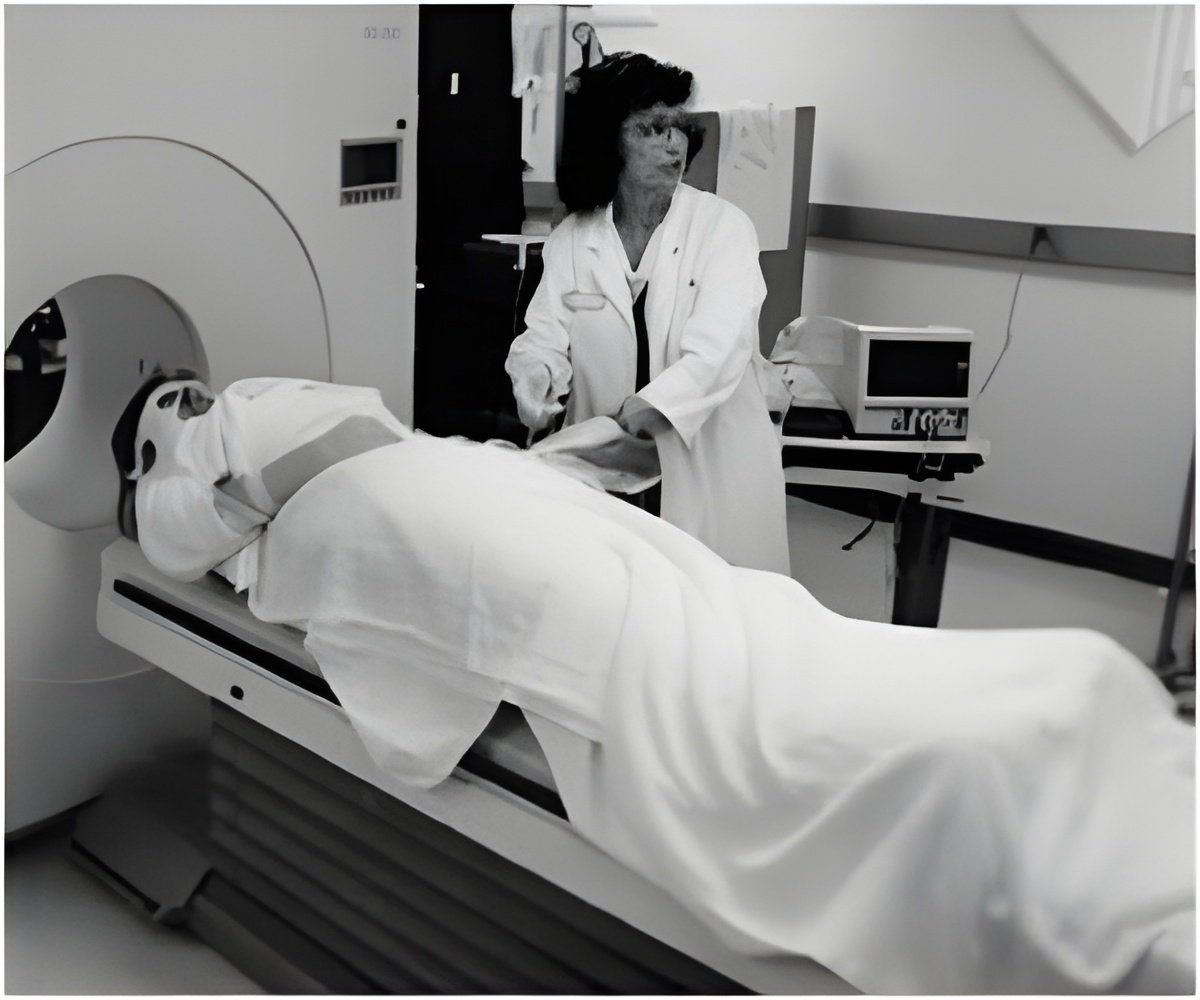
The study was published in the Feb. 22, 2013 issue of the journal PLOS ONE.
Neil Hayes, MD, MPH, associate professor of medicine and senior author, says, "Cancer is a disease caused by alteration in the DNA and RNA molecules of tumors. A cancer results when broken molecules initiate a cascade of abnormal signals that ultimately results in abnormal growth and spread of tissues that should be under tight control within the body."
"However, most common tumors, including head and neck cancer, have relatively little information in the public record as to how these signals coordinate to create different patterns of abnormalities. This study is among the largest ever published to document reproducible molecular tumor subtypes. Subtypes, such as those we describe, represent attractive models to understand and attack cancers for treatment and prognosis."
Dr. Hayes is a member of UNC Lineberger Comprehensive Cancer Center and national co-chair of the Data Analysis Sub-Group for The Cancer Genome Atlas, a program of the National Institutes of Health.
The team, composed of investigators from UNC and five other institutions, analyzed a set of nearly 140 HNSCC samples. By searching for recurrent patterns known as gene expression signatures, they were able to detect four gene expression subtypes. The subtypes are termed basal, mesenchymal, atypical, and classical based on similarities to established gene expression subtypes in other tumor types and expression patterns of specific genes.
 MEDINDIA
MEDINDIA

 Email
Email