Researchers are looking into the possibility of using white blood cell variation as a diagnostic, predictive and research tool in study of non-blood cancers.
Koestler is working in the Quantitative Biomedical Sciences program with Professors Margaret Karagas and Jason Moore. His focus is the development of computational and statistical tools for investigating the process of DNA methylation.
In methylation, a molecule known as a methyl group (chemically CH3—three hydrogen atoms and one carbon) attaches itself to the DNA. When this occurs, the DNA function can change dramatically. An example might be the methyl group blocking the expression of a tumor-suppressing gene.
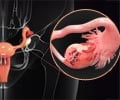
Koestler is the first author on a paper with Karagas and a host of colleagues from Dartmouth, Brown University, Oregon State University, the University of Minnesota and the University of California. Its subject is methylation in leukocytes (white blood cells) and their association with cancer in tissues and organs other than blood, such as bladder or ovarian cancers.
"When we have an illness or a disease, that does something to our immune system," Koestler explains. "It responds by providing whatever cells are necessary to combat that threat. In the blood, the leukocytes supply that immune response."
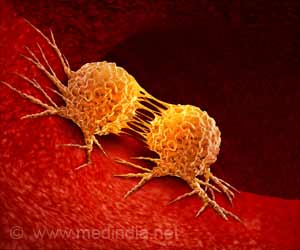
Methylation has been studied in biopsied cells from cancer patients, in comparison to cells of cancer-free individuals. "Those studies have compellingly shown there are very striking differences in methylation patterns between cancer and cancer-free subjects," Koestler says. "This brought us to also look at patterns of methylation in blood."
Advertisement
Using data from studies of ovarian, bladder, and head and neck cancers, the researchers demonstrated statistically significant correlations between the specific cancers and the methylation signatures that characterize leukocyte subsets.
Advertisement
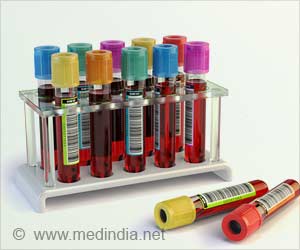
Analyzing the relative proportions of the leukocyte types in the blood sample can help predict the onset of a particular cancer or identify and diagnose a cancer in progress. The alternative of sampling a patient's blood is far preferable to undergoing an invasive surgical biopsy.
The advantages of using methylation patterns to assess proportions of white blood cell subtypes in cancer research extend beyond the bedside to the lab bench. Archival blood samples frozen and stored at some time in the past can now be used as research material, whereas existing methods typically require fresh blood samples with intact cells to assess white blood cell subtypes.
Source-Eurekalert