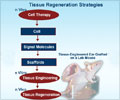
Northwestern University scientists have developed a powerful analytical method to direct stem cell differentiation.
Those in the fields of chemistry, materials engineering and nanotechnology could use this invaluable tool to assess which chemical and physical structures -- including size, shape and composition -- work best for a desired process or function.
Nanocombinatorics holds promise for screening catalysts for energy conversion, understanding properties conferred by nanostructures, identifying active molecules for drug discovery or even optimizing materials for tissue regeneration, among other applications.
Details of the method and proof of concept is published in the Proceedings of the National Academy of Sciences.
"With further development, researchers might be able to use this approach to prepare cells of any lineage on command," said Chad A. Mirkin, who led the work. "Insight into such a process is important for understanding cancer development and for developing novel cancer treatment methodologies."
Mirkin is the George B. Rathmann Professor of Chemistry in the Weinberg College of Arts and Sciences and professor of medicine, chemical and biological engineering, biomedical engineering and materials science and engineering. He also is the director of Northwestern's International Institute for Nanotechnology (IIN).
In this work, the researchers used molecules that bind proteins found in the natural cell environment, such as fibronectin, which could then be attached onto a substrate in various patterns. (Fibronectin is a protein that mediates cell adhesion.) The team rapidly prepared millions of textured features over a large area, which they call a library. The library consisted of approximately 10,000 fibronectin patterns having as many as 25 million features ranging in size from a couple hundred nanometers to several micrometers.
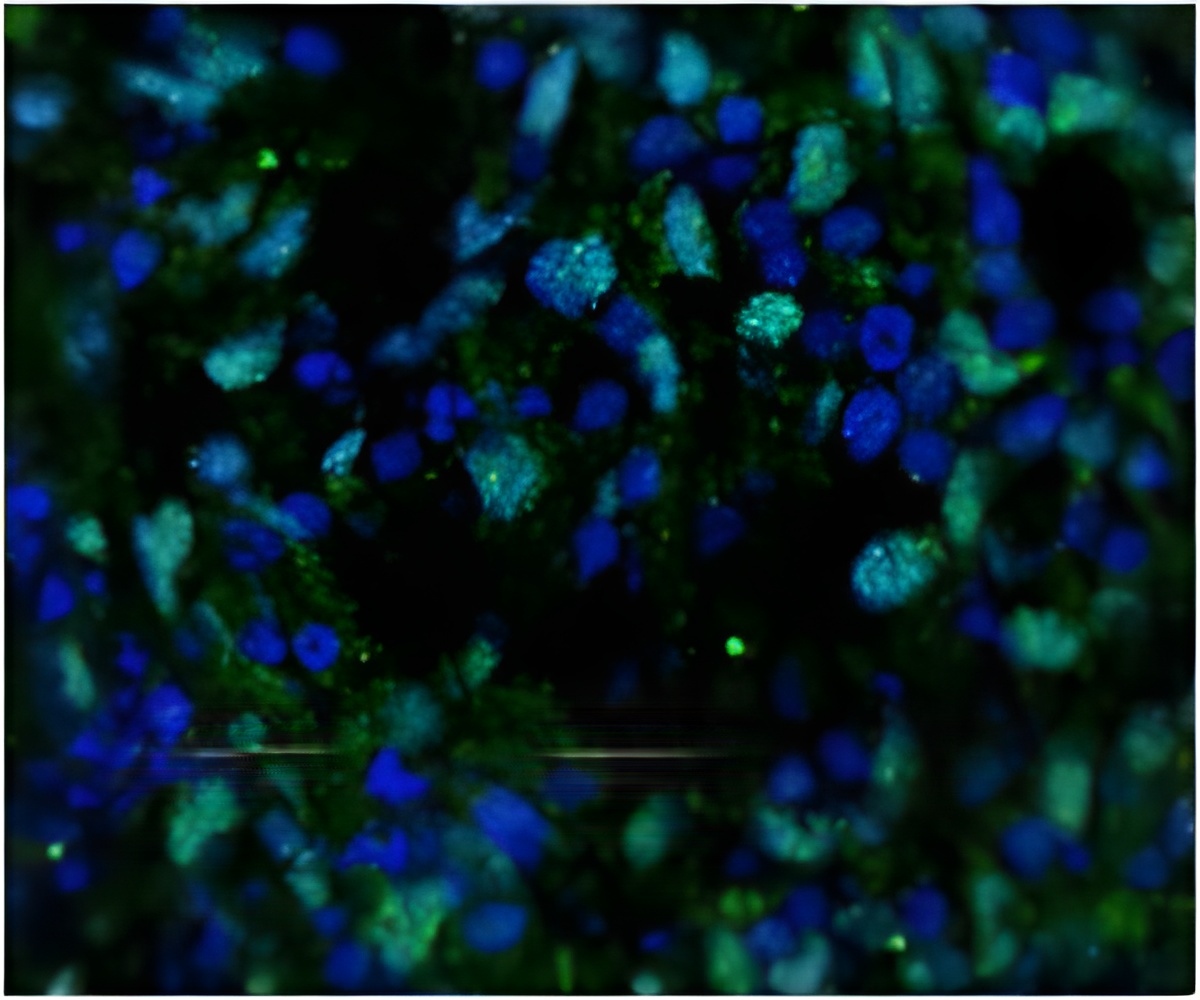
The researchers then introduced mesenchymal stem cells, or MSCs, to the library of millions of fibronectin features. (MSCs are multipotent stem cells that can differentiate into a variety of other cell types.)
"We let the cells sample the library and watched what happened," Mirkin said.
He and his team found areas with stem cell differentiation and areas with none. Nanoscale features, particularly protein spots that were 300 nanometers in diameter, were more likely to lead to bone-like cells than larger micron-scale features.
The researchers next built a library made up of only 300-nanometer dots and introduced stem cells. Almost all of the cells became bone-like.
"We want to make stem cells go down a predetermined path -- to make bone cells instead of nerve or muscle cells," Mirkin said. "Starting with millions of possibilities, we quickly zeroed in on the pattern of protein features that best directed the cells to become osteocytes."
This stem cell differentiation was accomplished without the use of additional chemical cues (beyond the proteins in the patterns). The transition from stem cell to osteocyte was dictated solely by the physical cues of the patterned structures. And the researchers demonstrated better control over stem cell differentiation than chemical reagent methods currently used.
"It doesn't stop with stem cells," Mirkin said. "Scientists can use nanocombinatorics to build libraries of structures that vary in shape, size and distance between particles and determine the best structures for controlling important events, like speeding up a catalytic reaction."
Source-Eurekalert
 MEDINDIA
MEDINDIA


 Email
Email