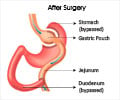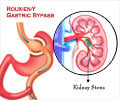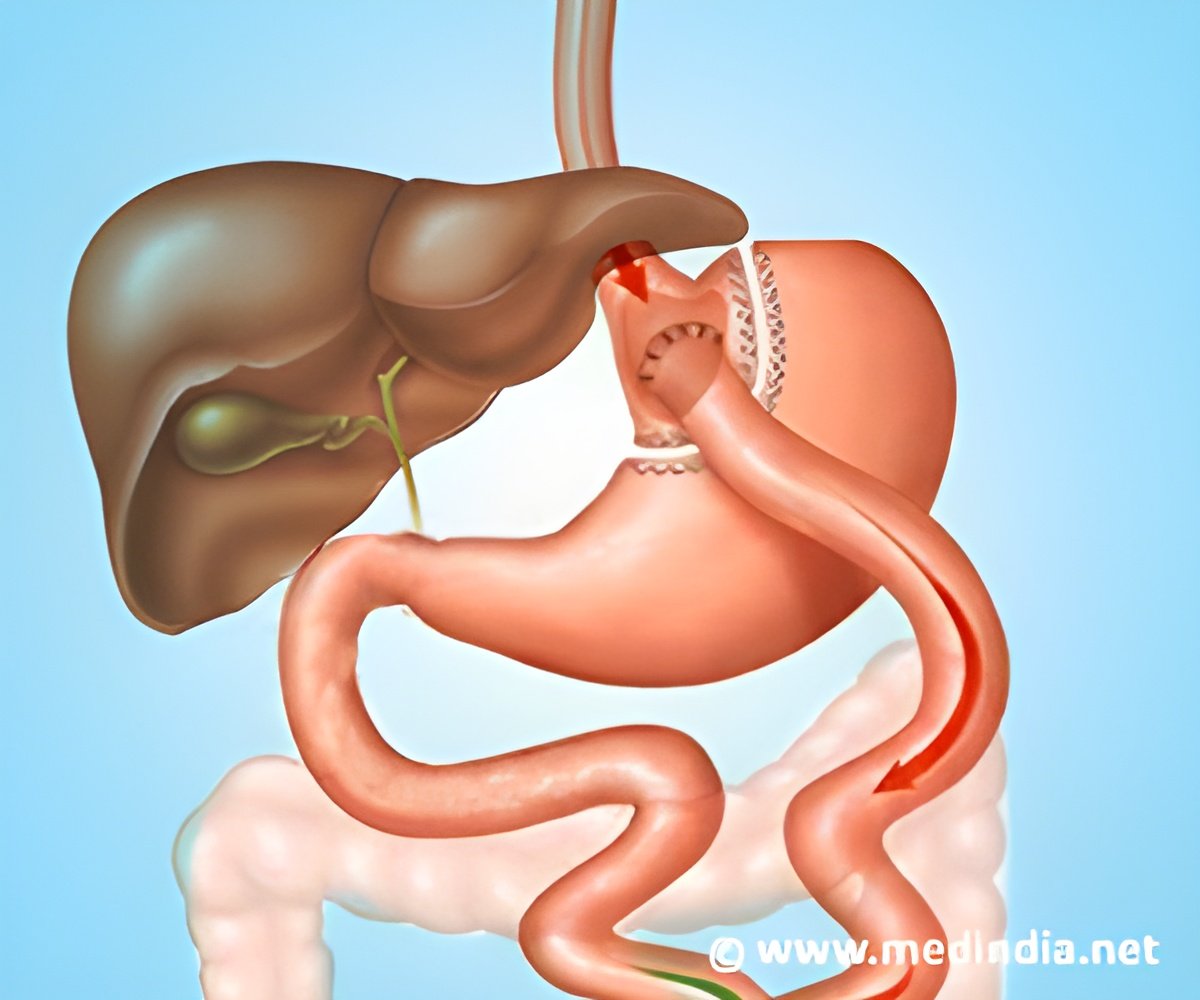
‘The therapeutic effects of gastric bypass provides a way in discovering novel drugs for obesity, type II diabetes and other metabolic diseases.’





The study was conducted by a team of researchers at the Massachusetts General Hospital Center for Engineering in Medicine (MGH-CEM) and Shriners Hospital for Children. Their report on the impact of the surgery on liver metabolism appeared last month in the journal Technology."The therapeutic effects of gastric bypass give us hope that we can discover novel drugs for obesity and diabetes" says Nima Saeidi, PhD, of the MGH-CEM, principal investigator on the project.
"Our goal is to reverse engineer the weight loss and resolution of blood sugar control without having to perform such an invasive procedure. To achieve this, we must first be able to capture how metabolic organs such as the liver, muscle, adipose tissue and even the small intestine reprogram themselves post-surgery to trigger the weight loss and reversal of insulin resistance"
To tackle this challenge, the team has been exploiting rapid advancements in mass spectrometry to effectively quantify thousands of biomolecules, namely metabolites and proteins, in order to obtain clues about how specific tissues or whole organs respond to gastric bypass.
In the present study, they utilized the state-of-the art SWATH proteomics workflow to discover approximately 100 proteins in rat livers, levels of which were significantly affected by Roux-en-Y gastric bypass (RYGB) surgery.
Advertisement
Having demonstrated the utility of this platform, the team plans to expand the analysis to study the effects of RYGB on multi-organ cross-talk via proteins secreted into the bloodstream.
Advertisement
He adds, "Going forward, developing network-based algorithms to handle time-series ’omics’ data will be imperative for unraveling complex cause-and-effect relationships, such as the impact of post-surgery plasma hormone levels on tissue transcriptional reprogramming."
Martin Yarmush, MD, PhD, director of the MGH-CEM and co-corresponding author of the paper, stresses the multi-disciplinary nature of the work.
"Investigators in the field have long sought after this type of multi-omic analysis on a rodent gastric bypass model to study metabolic disease, but the work required expertise in rodent surgery, multi-omic analytics, and computational biology. At the CEM, we enable engineering scientists to tackle complex problems in biomedicine at a world-class research hospital in collaboration with clinicians. This work will likely be of great interest to biotechnology companies aspiring to build their drug pipeline in the metabolic disease area."
Source-Eurekalert