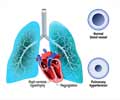
Over 6.3 million unique single nucleotide polymorphisms (SNPs) were identified in the genotypes of the patients with COPD, which is a progressive breathing disorder that limits airflow in the lungs. The genotypes of over 700 African Americans with COPD also were analyzed.
"Identifying single nucleotide polymorphisms associated with bronchodilator responsiveness may reveal genetic pathways associated with the pathogenesis of COPD and may identify novel treatment methods," said Megan Hardin, M.D., Instructor of Medicine at Harvard Medical School and researcher in the Channing Division of Network Medicine at Brigham and Women's Hospital, Boston.
Dr. Hardin, who presented the research, added that multiple genetic determinants likely influence bronchodilator responsiveness. Functional analysis of the SNPs will be conducted, she added.
"As we continue to analyze the data, we expect to identify other important SNPs," said Craig P. Hersh, M.D., who headed the study and is Assistant Professor, Harvard Medical School, and faculty member in the Channing Division of Network Medicine at Brigham and Women's Hospital.
All of the subjects studied had significant histories of smoking, with most (4,561), over 10 pack-years. All patients were genotyped, and their lung function was tested by spirometry before and after they used the bronchodilator medication albuterol, which relaxes muscles in the airways and increases airflow to the lungs. Spirometry measures the volume and flow of air that is exhaled.
Advertisement
In the presentation today, Dr. Hardin reported the top SNPs that thus far have been associated with each BDR outcome, but emphasized that additional analysis may reveal other SNPs with equally or greater influence on COPD patients' response.
Advertisement
Source-Eurekalert