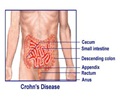
Multiple interacting mechanisms, including alterations in the intestinal microbiota, are suspected to lie behind irritable bowel syndrome (IBS), which is a common gastrointestinal functional disorder that can greatly affect the patient's well being.
A research article to be published on December 21, 2009 in the World Journal of Gastroenterology addresses this question. The research team from Finland quantified fourteen bacterial phylotypes which corresponded with bacterial species of the gastrointestinal (GI) tract from faecal samples of IBS patients and healthy controls. In their study, assays for analyzing phylotype specific bacterial alterations in association to IBS were developed and applied.They found a phylotype with 85% similarity to C. thermosuccinogenes was quantified in significantly different quantities among the diarrhoea-predominant (IBS-D) and control subjects, IBS-D and mixed symptom-subtype (IBS-M) subjects. A phylotype with 94% similarity to R. torques was more prevalent in IBS-D patients' intestinal microbiota than in that of control subjects. A phylotype with 93% similarity to R. torques was associated with control samples when compared with IBS-M. Additionally, a R. bromii-like phylotype was associated with constipation-predominant (IBS-C) patients in comparison to control subjects.
The researchers drew a conclusion that the detected altering phylotypes might be useful as targets in diagnostic, therapeutic and host-microbe interaction studies.
Source-Eurekalert
RAS