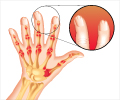
- Epigenetic landscape of rheumatoid arthritis (RA), a chronic inflammatory disorder decoded
- Connection between RA and Huntington’s disease, a fatal genetic disorder discovered
- The unexpected connection between RA and Huntington's disease opens up possibilities of discovering new therapeutic targets and drugs for both conditions.
The study is published in Nature Communications.
"We did not expect to find an overlap between rheumatoid arthritis and Huntington's disease but discovering the unexpected was the reason that we developed this technology. Now that we have uncovered this connection, we hope that it opens a door for treatment options for people living with either disease," said senior author Gary S. Firestein, MD, dean and associate vice chancellor of translational medicine at UC San Diego School of Medicine.
What is Epigenetics?
Epigenetics is the study of processes that alter the gene structure other than changes in the DNA sequence. The word “epigenetics” literally means in addition to changes in genetic sequence. Epigenetics can lead to modifications that are natural and essential to human growth and development; they change throughout people's lives. But if they occur improperly, there can be major adverse health and behavioral effects. Among several types of epigenetic processes that have been identified like methylation, acetylation, phosphorylation, ubiquitylation, and sumoylation, the best-known ones are DNA methylation, and chromatin modification, the former having been observed in cancer and many other illnesses.Epigenetic mechanisms have been implicated so far in cancers of almost all types, cognitive dysfunction, and respiratory, cardiovascular, reproductive, autoimmune, and neuro-behavioral illnesses.
The suspected drivers of the changes include a variety of environmental factors (including heavy metals, pesticides, diesel exhaust, tobacco smoke, polycyclic aromatic hydrocarbons, stress, radioactivity, virus, bacteria, hormones), basic nutrients and lifestyle choices.
Study
The team studied the epigenome in cells from the joints of patients with RA and compared it with a group of patients with osteoarthritis (OA), a cartilage degeneration disease.The research team involved developed a novel algorithm, or set of computational rules, called EpiSig, which integrated diverse epigenomic data subsets which are usually difficult to be analyzed together and made them into a single analysis.
The platform clustered regions with similar epigenomic profiles that shared common functionality across all RA and OA. This reduced the number of epigenetic combinations in the genes of patients with RA and helped them identify new cell signaling pathways.
The team analyzed the data sets through an elaborate process that examines modifications in DNA, histone and chromatin.
A huge amount of data produced (12 trillion bytes) were then analyzed using EpiSig.
This methodology can also be used to find connections between diseases other than rheumatoid arthritis too.
Significance of the findings and future of EpiSig
Treatment of RA includes non-steroidal anti-inflammatory drugs (NSAIDs) that relieve pain and reduce inflammation, corticosteroid medications and disease-modifying antirheumatic drugs (DMARDs) that are long-term medications that slow the progression of rheumatoid arthritis by stopping the immune system from attacking healthy tissue.Biologics are a type of DMARD that specifically target steps in the inflammatory process; but, even with targeted therapies, it is difficult to obtain remission of RA.
Nowadays, most efforts to develop new therapies rely on candidate gene approaches where the potential to discover entirely new and unanticipated pathogenic processes that contribute to disease is limited.
EpiSig, the new methodology and dataset have provided a new way to identify RA-specific targets that can be used to develop novel therapeutic agents.
EpiSig is a genome-wide unbiased methodology that can identify surprising pathways and genes involved in immune dysfunction. Integration of multiple epigenetic technologies will make it easier to define the pathogenesis of disease and discover non-obvious targets.
The methodology can be applied to any immune-mediated disease if the datasets are available.
EpiSig can be used in the future to validate a large number of unanticipated targets and pathways, to explore the diversity between patients and to individualize treatment.
Rheumatoid Arthritis (RA)
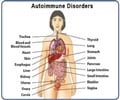
RA is an autoimmune disease that causes chronic abnormal inflammation, primarily in the joints. Being an autoimmune disease where the body’s immune system attacks its own tissues and organs, other organs like the heart and blood vessels can also be affected.Rheumatoid arthritis presumably results from a combination of genetic and many unknown environmental factors; the most significant genetic risk factors are variations in the human leukocyte antigen (HLA) genes. HLA genes code for proteins that help the immune system distinguish the body's own proteins from proteins made by foreign invaders like viruses and bacteria.
References:
- Rizi Ai, Teresina Laragione, Deepa Hammaker, David L. Boyle, Andre Wildberg, Keisuke Maeshima, Emanuele Palescandolo, Vinod Krishna, David Pocalyko, John W. Whitaker, Yuchen Bai, Sunil Nagpal, Kurtis E. Bachman, Richard I. Ainsworth, Mengchi Wang, Bo Ding, Percio S. Gulko, Wei Wang & Gary S. Firestein. "Comprehensive epigenetic landscape of rheumatoid arthritis fibroblast-like synoviocytes." Nature Communications (2018) doi:10.1038/s41467-018-04310-9
- Weinhold B. Epigenetics: The Science of Change. Environmental Health Perspectives. (2006); 114(3):A160-A167.
Source-Medindia