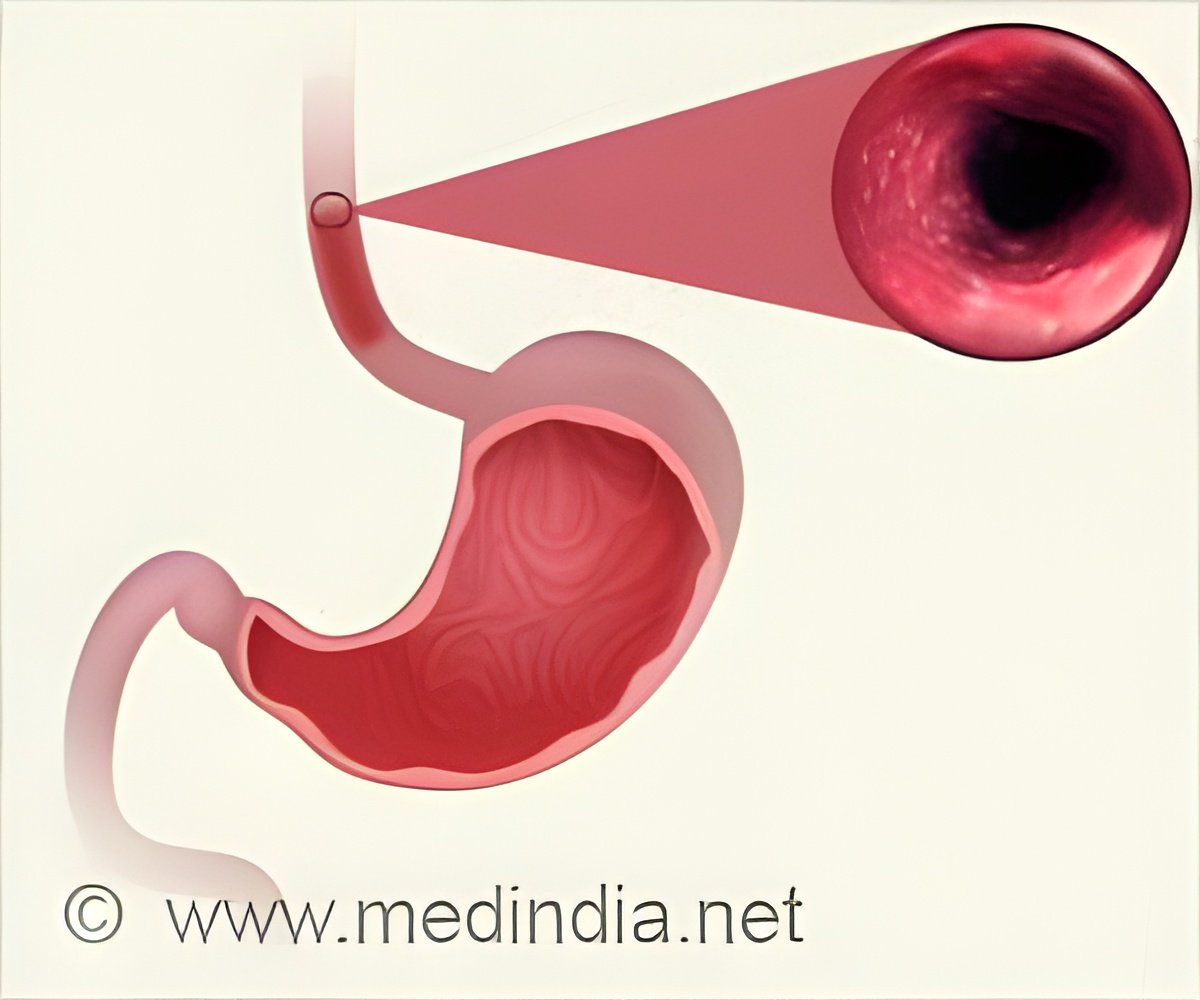
‘Barrett's Esophagus (BE) is a risk factor for developing Esophageal Adenocarcinoma (EAC), a deadly, highly intractable cancer, but only a tiny proportion of BE patients will progress to cancer.’
Tweet it Now
A small number of BE patients--just .2 percent per year--will go on to develop a highly lethal, treatment-resistant cancer, known as Esophageal Adenocarcinoma (EAC). Despite advances in therapy, prospects for EAC patients remain bleak--fewer than 15 percent survive beyond 5 years. (While the incidence of EAC in the United States has remained low, it has risen alarmingly in recent years.) Understanding why most BE patients avoid this fate and some don't has been a challenge for the medical community. Evolutionary oncologists like Carlo Maley, a researcher at the Biodesign Center for Personalized Diagnostics and a senior author of the new study, believe a better understanding of the evolutionary dynamics of this process may hold the key. The study examines these dynamics over at least 6 years of surveillance. Of the 8 BE patients examined, 4 remain stable and 4 progress to cancer.
"By taking a host of minute samples across the surface of the esophagus, and across many years while these patients were under surveillance to detect cancer, we had an unprecedented view of the dynamics of carcinogenesis," Maley says.
Sensing the threat
BE presents a conundrum for clinicians. Because no reliable means exist at present to distinguish progressors from non-progressors, the only cautionary measures available involve repeated surveillance of BE esophageal cells through endoscopy and biopsy to try to catch EAC-linked abnormalities at an early stage, or methods to burn away the lining of the esophagus.
Advertisement
As the authors note, better predictions will rely in part on testing BE samples at multiple points in time, and an examination of cells extracted from different locations in the esophageal tissue. One positive consequence of aggressive BE cell surveillance is that it has provided researchers with a rich library of data that can be mined using new methods in order to tease out critical factors governing progression vs non-progression to cancer.
Advertisement
Barrett's Esophagus can develop over time when digestive acid backs up from the stomach into the esophagus, causing damage and growth of precancerous cells. To accurately assess the evolutionary dynamics involved in progression to cancer, researchers need more fine-grained analyses of BE cells, to tease out details that may not be detectable in whole biopsies containing millions of cells.
In BE, the columnar architecture of the epithelium is similar to that of the intestine. Here, well-like structures in the tissue known as crypts appear. At the base of those crypts are stem cells, which replenish the epithelium as older cells migrate up the sides of the crypts and then die off. The study represents the first genome-wide analysis of the evolutionary dynamics in BE at the level of individual crypts. (While crypts may contain more than a single stem cell, these cells tend to be genetically uniform. For this reason, crypt sampling is closely analogous to sampling single BE cells.)
Researchers would like to know why most cases of BE appear to be evolutionarily static over time. Either genetic alterations tend to be rare or, if they are common, cells carrying those alterations do not expand to levels detectable through conventional biopsy. The new study examines the genomes of single crypts in BE to take a closer look at when and where genetic variants arise and how the evolutionary process plays out.
Ominous progression
Advancement from BE to EAC is a process characterized by mounting genomic instability. Over time, BE patients can accumulate mutations in their esophageal lesions, altering the genetic makeup of these cells. Such genetic variation is regarded as a valuable predictor of progression to cancer, though the highly complex dynamics of this process are still being worked out.
The study focused on losses and gains of large regions of chromosomes in the BE cells. Aberrations in the chromosomes are high in those who go on to develop EAC, even 4 years before progression and remain low in non-progressors, pointing to the value of chromosomal diversity as a diagnostic indicator.
The study also examined a phenomenon known as genome doubling. This results from faulty cell division, which creates a cell with twice the normal number of chromosomes. Those likely to progress to EAC were also more likely to experience genome doubling, which is presaged by an increasing rate of accumulation of chromosomal aberrations.
The study examines genetic variation in BE crypts, comparing these with the variation found through examination of biopsies. Multiple biopsies and crypt samples were examined from 8 BE patients at two different time points. Four of these patients progressed to EAC and 4 did not.
Results comparing biopsy and single crypt information show that genetic alterations are indeed rare, even at the crypt level, and that Barrett's lesions tend to arise from a single ancestral cell gone awry. Further, the study selects cells from different regions of the esophagus and finds that genetic alterations were more common in samples taken near the base of the esophagus, known as the gastro-esophageal junction.
New findings
The study addresses five previously unanswered questions concerning BE. As the authors stress, the results offer new insights into the general process of cell progression to cancer, which may be applicable across many, if not all forms of the disease.
Results indicate that
BE tissue in most cases arises from a single altered ancestral cell
expansion of cancerous clones (identical cells of common ancestry) is rare
cells sampled near the gastro-esophageal junction accumulate more genomic alterations than those found in other regions of the esophagus, making them better targets for diagnosis
there are dramatic changes in mutation rate during progression and these may occur early in the process of cancer progression and
genetic diversity as measured through biopsy in Barrett's patients is comparable to that observes in individual crypts, indicating that biopsy analysis is adequate for assessing the evolutionary characteristics of BE cells and their likelihood of progression.
Continued work in this area promises to untangle the complex network of evolutionary factors at play when cells are directed away from their normal course and toward the fateful path of cancer.
Source-Eurekalert