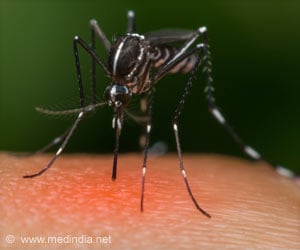
The protozoan cannot survive without this enzyme, but even though the enzyme has many look-alikes in other organisms, people do not make it.
Together these characteristics make the enzyme an ideal target for new anti-malarial drugs.
The protein's structure might have remained an enigma, had it not been the "unreasonable optimism" of Joseph Jez, PhD, associate professor of biology in Arts and Sciences, which carried his team through a six-year-long obstacle course of failures and setbacks.
"What my lab does is crystallize proteins so that we can see what they look like in three dimensions," Jez said.
"The idea is that if we know a protein's structure, it will be easier to design chemicals that would target the protein's active site and shut it down," he added.
Advertisement
The research was published in the January 6 issue of the Journal of Biological Chemistry (JBC).
Advertisement