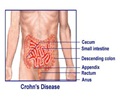
Researchers at University of Cincinnati (UC) have identified a new biomarker that could help predict a person's risk of developing colon cancer and how aggressive it may become.
The team has identified 'hotspots', areas of deleted genetic data, that play a key role in regulating gene expression and influence colon cancer progression.Researchers speculate that these hotspots could be used as a biomarker for colon cancer.
This study is the first to explain how a gene-AMACR-is regulated in relation to cancer development and to identify specific genetic events (a polymorphism and somatic cell mutations) related to colon cancer.
For this study, the research team looked at the actions of the AMACR gene in human tissue.
AMACR breaks down branched-chain fatty acids, a type of molecule only found in animals that eat plants.
Previous study showed that plant-derived fatty acids, such as those found in red meat and dairy products, can accelerate cancer growth.
Advertisement
The researchers analyzed the AMACR gene's abnormal expression patterns using a sophisticated laser-capture microdissection technique to identify the key biological events that lead to colon cancer progression.
Advertisement
Besides discovering the hotspots that trigger abnormal AMACR expression, they also identified specific proteins (transcription factors) that would normally bind to the deleted sequences to maintain normal gene expression.
"Our hope is that this new knowledge will help us develop better diagnostic tools for colon cancer," said Zhang.
The study is published in the Jan. 16, 2009, issue of PLoS Genetics.
Source-ANI
PRI/L