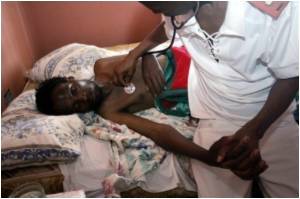
Some of the structures solved by the centers come from well-known, headline-grabbing organisms, like the H1N1 flu virus.
The two leading canters are The Center for Structural Genomics of Infectious Diseases (CSGID), led by Wayne Anderson and the Seattle Structural Genomics Center for Infectious Disease (SSGCID), led by Peter Myler.
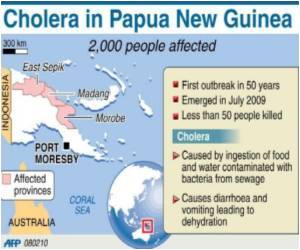
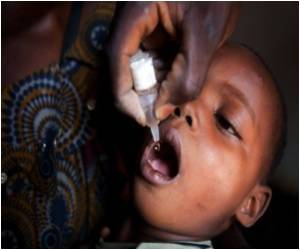
The Centers' mission is to apply genome-scale approaches in solving protein structures from biodefense organisms, as well as those causing emerging and re-emerging diseases.
Other structures solved come from little known or emerging pathogens that cause disease and death, but have been less well studied by the research community.
Recently, scientists at CSGID determined the structure of a crucial enzyme in the shikimate pathway of Clostridium difficile, which is the most serious cause of antibiotic-associated diarrhea in humans.
Advertisement
The structures solved by the Centers are immediately made available to the international scientific community through the NIH-supported Protein Data Bank (www.pdb.org), providing a "blueprint" for development of new drugs, vaccines and diagnostics.
Advertisement