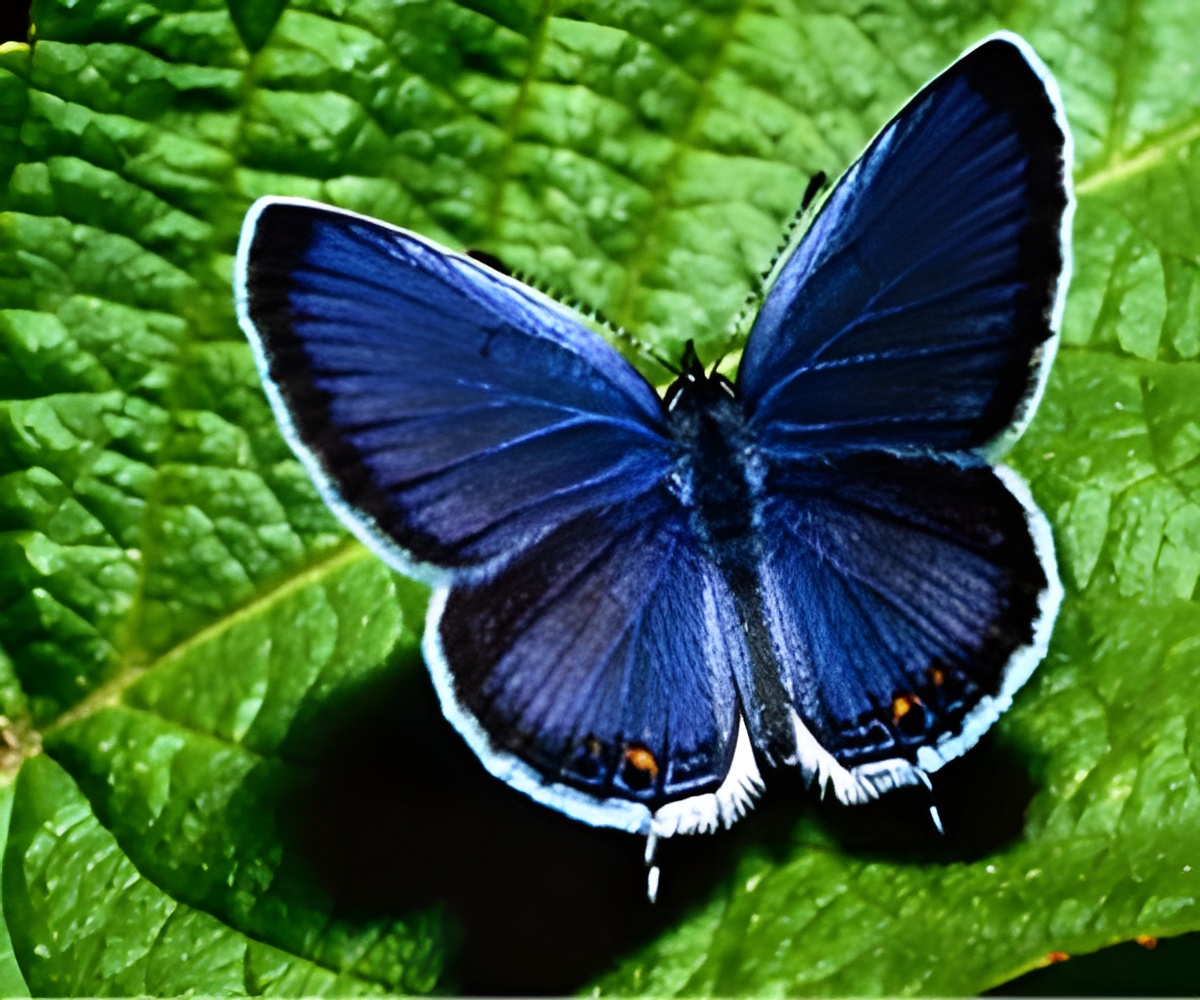
The red postman is an abundant tropical butterfly found in Central and South America.
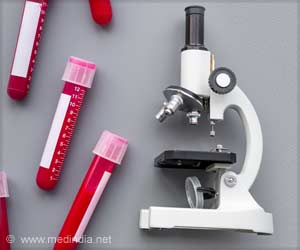
The results showed the internal bacterial diversity of the red postman was halved when it morphed from the caterpillar to the chrysalis, or pupal stage, then doubled after the pupae turned into active adult butterflies.
The butterflies were collected at a field site in Gamboa, Panama, and data analysis was done at CU-Boulder.
The main motivation for choosing the red postman for study is that its genus is the only one that feeds on pollen, a rich source of amino acids, Hammer said. Most butterflies feed on nectar-essentially sugar water-leading to abbreviated lifespans lasting only days or weeks.
But the red postman apparently has found a way to digest and then extract nutrients from pollen grains, likely expanding its lifespan by several months.
Advertisement
The subject has been published online in the journal PLOS ONE.
Advertisement