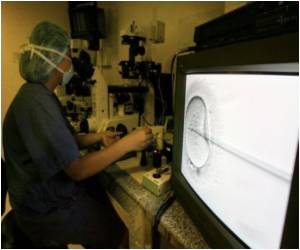
Data from a subanalysis of the phase III GIST-Regorafenib In Progressive Disease (GRID) trial indicated that this blood-based screening technology may provide physicians with a real-time, comprehensive picture of a patient's tumor mutations, according to George D. Demetri, M.D., director of the Ludwig Center at Dana-Farber Cancer Institute and Harvard Medical School in Boston, Mass.
"Our results show that it is possible to sum the total of all of the heterogeneity in a cancer and get a clear picture of the entire tumor burden, using a simple blood sample," Demetri said.
In this era of targeted cancer therapies, the goal is to focus cancer treatments on a specific molecular target. However, as researchers discover more about cancers and their heterogeneity, they are finding many patients have anywhere from one to dozens of different mutations in their tumors.
"It is a real issue that when you do a biopsy on one tumor, and then biopsy a different tumor in that same patient a few inches away or on the other side of the body, you may get a different answer when you do the molecular analysis," Demetri said. "With this blood test, you get a robust summary statement about all the different mutations present across the different tumors in the body. I believe this testing technology has promise to become a standard part of care in the next five to 10 years."
Data from the main analysis of the phase III GRID study showed that the molecularly targeted drug regorafenib significantly improved progression-free survival compared with placebo for patients with GIST. The researchers hope these results will ultimately lead to the drug's approval by the U.S. Food and Drug Administration (FDA), according to Demetri. The drug is intended to treat patients with advanced GIST whose disease has failed to be controlled by the only two other FDA-approved therapies for GIST, imatinib and sunitinib (Sutent).
Advertisement
Mutations in the KIT gene were detected in 60 percent of the blood samples compared with 65 percent of the tumor tissue samples. However, when focusing their analysis on secondary KIT mutations, which are the mutations that drive resistance to targeted therapies like imatinib and sunitinib, the researchers found mutations in 48 percent of blood samples compared with only 12 percent of tissue samples. In addition, nearly half of blood samples in which secondary KIT mutations were found harbored multiple secondary mutations.
Advertisement
According to Demetri, the results show a clear association between the presence of different cancer-driving gene mutations in patients' blood samples and clinical outcomes.
"By using this technology, we hope to develop the most rational drug combinations and better tests to match patients with the most effective therapies going forward," Demetri said.
Press registration for the AACR Annual Meeting 2013 is free to qualified journalists and public information officers: www.aacr.org/PressRegistration.
About the American Association for Cancer Research
Founded in 1907, the American Association for Cancer Research (AACR) is the world's first and largest professional organization dedicated to advancing cancer research and its mission to prevent and cure cancer. AACR membership includes more than 34,000 laboratory, translational and clinical researchers; population scientists; other health care professionals; and cancer advocates residing in more than 90 countries. The AACR marshals the full spectrum of expertise of the cancer community to accelerate progress in the prevention, biology, diagnosis and treatment of cancer by annually convening more than 20 conferences and educational workshops, the largest of which is the AACR Annual Meeting with more than 17,000 attendees. In addition, the AACR publishes eight peer-reviewed scientific journals and a magazine for cancer survivors, patients and their caregivers. The AACR funds meritorious research directly as well as in cooperation with numerous cancer organizations. As the scientific partner of Stand Up To Cancer, the AACR provides expert peer review, grants administration and scientific oversight of team science and individual grants in cancer research that have the potential for near-term patient benefit. The AACR actively communicates with legislators and policymakers about the value of cancer research and related biomedical science in saving lives from cancer. For more information about the AACR, visit www.AACR.org.
Abstract Number: LB-295
Presenter: George D. Demetri, M.D.
Title: Detection of oncogenic kinase mutations in circulating plasma DNA and correlation with clinical benefit in the phase III GRID study of regorafenib vs placebo in TKI-refractory metastatic GIST
Authors: George D. Demetri 1, Michael Jeffers 2, Peter Reichardt 3, Yoon-Koo Kang 4, Jean-Yves Blay 5, Piotr Rutkowski 6, Hans Gelderblom 7, Peter Hohenberger 8, Michael Leahy 9, Margaret von Mehren 10, Heikki Joensuu 11, Giuseppe Badalamenti 12, Martin Blackstein 13, Axel Le Cesne 14, Patrick Schffski 15, Robert G Maki 16, Sebastian Bauer 17, Binh Bui Nguyen 18, Jianming Xu 19, Toshirou Nishida 20, John Chung 21, Chetan D. Lathia 22, Christian Kappeler 23, Iris Kuss 23, Dirk Laurent 23, Paolo G Casali 24. 1Dana-Farber Cancer Inst., Boston, MA; 2 Bayer HealthCare, Montville, NJ; 3Department of Hematology, Oncology and Palliative Medicine, HELIOS Klinikum, Bad Saarow, Germany; 4Department of Oncology, University of Ulsan College of Medicine, Asan Medical Center, Seoul, Korea, Republic of; 5 Centre Leon Berard and University Claude Bernard, Leon, France; 6 Department of Soft Tissue/Bone Sarcoma and Melanoma, Maria Sktodowska-Curie Memorial Cancer Center and Institute of Oncology, Warsaw, Poland; 7 Department of Clinical Oncology, Leiden University Medical Centre, Leiden, Netherlands; 8 Div. of Surgical Oncology & Thoracic Surgery Mannheim University Medical Center, Mannheim, Germany; 9 The Christie NHS Foundation Trust, Manchester, United Kingdom; 10 Fox Chase Cancer Center, Philadelphia, PA; 11 Department of Oncology, Helsinki University Central Hospital, Helsinki, Finland; 12 Department of Oncology, University of Palermo, Palermo, Italy; 13 Medical Oncology Unit, Mount Sinai Hospital, Toronto, ON, Canada; 14 Department of Medicine, Gustave Roussy Institute, Paris, France; 15 Department of General Medical Oncology, University Hospitals Leuven, Leuven, Belgium; 16 Department of Medicine, Mount Sinai School of Medicine, New York, NY; 17 Sarcoma Center, West German Cancer Center, University of Duisburg-Essen, Essen, Germany; 18 Institut Bergoni©, Bordeaux, France; 19 Affiliated Hospital Cancer Center, Academy of Military Medical Sciences, Beijing, China; 20 Department of Surgery, Osaka Police Hospital., Osaka, Japan; 21 Bayer Healthcare Pharmaceuticals, Wayne, NJ; 22Alexion Pharmaceuticals, Cheshire, CT; 23 Bayer Pharma AG, Berlin, Germany; 24 Fondazione IRCCS Istituto Nazionale dei Tumori, Milan, Italy
Background: GRID is a phase III study for patients with advanced gastrointestinal stromal tumors (GIST) following failure of imatinib (I) and sunitinib (S) who were randomized to receive either the multikinase inhibitor regorafenib (R) or placebo (P). R demonstrated a highly significant improvement in progression-free survival compared with P (HR 0.27, p<0.0001). A preplanned retrospective biomarker analysis was conducted to assess GIST genotypes in GRID patients and to explore the possible impact of different driver oncogene mutations on clinical outcomes.
Methods: DNA was isolated from archival tumor tissue and analyzed for KIT mutations via Sanger sequencing. The expectation was that primary KIT mutations would be detectable in archival tissue but that secondary KIT mutations may be undetectable in tissues obtained before treatment with I or S. To overcome this potential limitation, plasma samples drawn at GRID study entry, post I and S failure, were used as a source of circulating DNA for evaluation of GIST oncogenic mutations (KIT, PDGFRA, BRAF) via BEAMing technology.
Results: KIT mutations were detected in 83 of 138 (60%) plasma samples and 64 of 99 (65%) tumor tissue samples analyzed. Primary KIT exon 11 and 9 mutations were identified in approximately 42% and 18% of the tissue samples, respectively. The frequency of the canonical exon 9 mutations was similar for plasma and tissue samples, showing consistency between mutation-detection technologies. With limitations of tumor-based assays, a lower incidence of secondary KIT resistance mutations was detected in patient-matched archival tumor tissue compared with plasma samples: resistance mutations were detected in 12% of tissue samples vs 48% of plasma samples. Most (76%) secondary KIT mutations detected in plasma DNA were located in the KIT activation loop encoding structural alterations known to mediate resistance to I and S. Nearly half of the plasma samples in which secondary KIT mutations were identified harbored multiple secondary mutations, consistent with the results of previous studies on fresh tumor biopsies taken following resistance to both I and S. R was clinically active compared with P in all KIT mutational subgroups evaluated (HR 0.27 in patients with KIT exon 9 mutations; HR 0.25 in patients with secondary KIT mutations identified via plasma DNA).
Conclusions: In GIST patients from the GRID trial, driver oncogenic mutations and secondary oncogenic mutations leading to I and S resistance are readily detectable via BEAMing of circulating DNA from plasma. BEAMing may provide a real-time assessment of tumor genotype in GIST and other cancers using blood-derived circulating DNA, that may be more comprehensive than tumor sampling. GIST patients with a wide spectrum of primary and secondary mutations in oncogenic kinases benefit from treatment with R.
Source-Newswise